Machine Learning
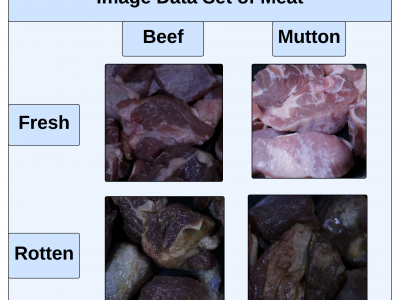
This paper presents an innovative Internet of Things (IoT) system that integrates gas sensors and a custom Convolutional Neural Network (CNN) to classify the freshness and species of beef and mutton in real time. The CNN, trained on 9,928 images, achieved 99% accuracy, outperforming models like ResNet-50, SVM, and KNN. The system uses three gas sensors (MQ135, MQ4, MQ136) to detect gases such as ammonia, methane, and hydrogen sulfide, which indicate meat spoilage.
- Categories:
 289 Views
289 ViewsThe TiHAN-V2X Dataset was collected in Hyderabad, India, across various Vehicle-to-Everything (V2X) communication types, including Vehicle-to-Vehicle (V2V), Vehicle-to-Infrastructure (V2I), Infrastructure-to-Vehicle (I2V), and Vehicle-to-Cloud (V2C). The dataset offers comprehensive data for evaluating communication performance under different environmental and road conditions, including urban, rural, and highway scenarios.
- Categories:
 518 Views
518 ViewsThe dataset provides crop-type surveys for Canada's prairie provinces (Manitoba, Saskatchewan and Alberta) in 2020 and 2021. The data were collected via windshield survey(driving through the countryside with GPS-enabled data collection software and satellite imagery). Crop-type points and their geographic coordinates on the ground were gathered using data collection software. Field boundaries were identified on satellite imagery. A single observation point is dropped in a homogeneous area within the field.
- Categories:
 246 Views
246 ViewsEndemic fish species are key components in seafood culinary excursions. Despite the increasing interest in leveraging technology to enhance various seafood culinary activities, there is a shortage of comprehensive datasets containing images of seafood used in artificial intelligence research, mainly those showcasing endemic fish. This research endeavors to bridge this gap by increasing the accuracy of fish recognition and introducing a new dataset comprising images of native fish for application in various machine-learning investigations.
- Categories:
 168 Views
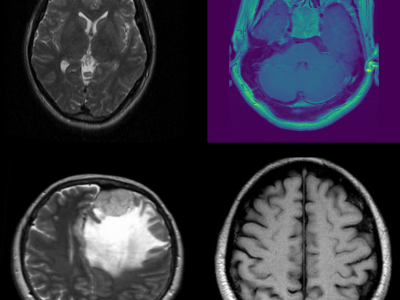
168 ViewsThis dataset consists of MRI images of brain tumors, specifically curated for tasks such as brain tumor classification and detection. The dataset includes a variety of tumor types, including gliomas, meningiomas, and glioblastomas, enabling multi-class classification. Each MRI scan is labeled with the corresponding tumor type, providing a comprehensive resource for developing and evaluating machine learning models for medical image analysis. The data can be used to train deep learning algorithms for brain tumor detection, aiding in early diagnosis and treatment planning.
- Categories:
 709 Views
709 ViewsTo download this dataset without purchasing an IEEE Dataport subscription, please visit: https://zenodo.org/records/13896353
Please cite the following paper when using this dataset:
- Categories:
 616 Views
616 Viewsyou can download these datasets from OpenML: https://www.openml.org/search?type=data&status=active&tags.tag=2019_mult...
- Categories:
 106 Views
106 Viewsyou can download these datasets from OpenML: https://www.openml.org/search?type=data&status=active&tags.tag=2019_mult...
- Categories:
 55 Views
55 Views
Liver cancer treatment, especially for metastatic cases, poses significant challenges in accurately targeting tumours while sparing healthy tissue. Radioembolisation with yttrium-90 (Y-90) microspheres is a promising technique, but precise imaging of microsphere distribution is crucial. This study utilises T-PEPT, a novel Positron Emission Particle Tracking (PEPT) algorithm that combines topological data analysis with machine learning to identify Y-90 microsphere clusters in a digital twin of a patient's liver.
- Categories:
 86 Views
86 ViewsThe Unified Multimodal Network Intrusion Detection System (UM-NIDS) dataset is a comprehensive, standardized dataset that integrates network flow data, packet payload information, and contextual features, making it highly suitable for machine learning-based intrusion detection models. This dataset addresses key limitations in existing NIDS datasets, such as inconsistent feature sets and the lack of payload or time-window-based contextual features.
- Categories:
 705 Views
705 Views