Biomedical and Health Sciences
This paper presents a dataset of brain Electroencephalogram (EEG) signals created when Malayalam vowels and consonants are spoken. The dataset was created by capturing EEG signals utilizing the OpenBCI Cyton device while a volunteer spoke Malayalam vowels and consonants. It includes recordings obtained from both sub-vocal and vocal. The creation of this dataset aims to support individuals who speak Malayalam and suffer from neurodegenerative diseases.
- Categories:
 2591 Views
2591 Views
This study presents a comprehensive dataset to analyze risk factors associated with cardiovascular disease. The dataset comprises various patient attributes, including gender, age, total cholesterol, HDL (high-density lipoprotein), triglycerides, non-HDL (non-high-density lipoprotein), NIH-Equ-2, and direct LDL (low-density lipoprotein). These attributes comprise 25,991 patient data, robustly representing a large population sample.
- Categories:
 584 Views
584 ViewsWe introduce an online-offline Iraquian hand-drawing dataset for early Parkinson’s disease detection, exclusively collected using smartphones, thus eliminating the need for specialized equipment like digitizing tablets and pens. Our dataset comprises data from 30 healthy individuals (17 men, 13 women) with an average age of 56 years (SD = 6.12) and 30 PD patients (23 men, 7 women) with an average age of 60 years (SD = 4.91), gathered at Marjan Hospital in Hilla, Babil Governorate, Iraq.
- Categories:
 584 Views
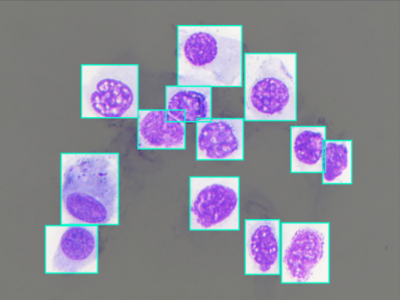
584 ViewsNasal Cytology, or Rhinology, is the subfield of otolaryngology, focused on the microscope observation of samples of the nasal mucosa, aimed to recognize cells of different types, to spot and diagnose ongoing pathologies. Such methodology can claim good accuracy in diagnosing rhinitis and infections, being very cheap and accessible without any instrument more complex than a microscope, even optical ones.
- Categories:
 807 Views
807 ViewsTo address the challenges faced by patients with neurodegenerative disorders, Brain-Computer Interface (BCI) solutions are being developed. However, many current datasets lack inclusion of languages spoken by patients, such as Telugu, which is spoken by over 90 million people in India. To bridge this gap, we have created a dataset comprising Electroencephalograph (EEG) signal samples of commonly used Telugu words. Using the Open-BCI Cyton device, EEG samples were captured from volunteers as they pronounced these words.
- Categories:
 461 Views
461 Views
The AnxiECG-PPG Database contains synchronized electrocardiogram (ECG) and mobile-acquired photoplethysmography (PPG) recordings from 47 healthy participants. Moreover, the acquisition protocol assesses three distinct states: a 5-minute Baseline, a 1-minute Physical Activated State, and a Psychological Activated state provoked through emotion-induced videos (negative, positive, and neutral emotion valence).
- Categories:
 1509 Views
1509 ViewsDespite advances in vascular replacement and repair, fabricating small-diameter vascular grafts with low thrombogenicity and appropriate tissue mechanics remains a challenge. A wide range of platforms have been developed to use plant-derived scaffolds for various applications. Unlike animal tissue, plants are primarily composed of cellulose which can offer a promising, nonthrombogenic alternative capable of promoting cell attachment and redirecting blood flow.
- Categories:
 194 Views
194 ViewsThis paper introduces a dataset capturing brain signals generated by the recognition of 100 Malayalam words, accompanied by their English translations. The dataset encompasses recordings acquired from both vocal and sub-vocal modalities for the Malayalam vocabulary. For the English equivalents, solely vocal signals were collected. This dataset is created to help Malayalam speaking patients with neuro-degenerative diseases.
- Categories:
 2857 Views
2857 Views
Normal
0
false
false
false
EN-US
X-NONE
AR-SA
- Categories:
 208 Views
208 ViewsTo access this dataset without purchasing an IEEE Dataport subscription, please visit: https://zenodo.org/doi/10.5281/zenodo.11711229
Please cite the following paper when using this dataset:
- Categories:
 981 Views
981 Views