Biomedical and Health Sciences

taken from a manta-ro experiment in five psychiatric patients after physical remediation, and the images were cut (documented by the Ethics Committee, WHO)
This dataset has sorted out formats that are suitable for classification and recognition using CNN.
By drawing a mandala pattern, participants can project their inner emotions, potential beliefs, and inner worlds
This is a magical adventure that crosses the boundaries of the mind. This article will delve into the background, methods, and possible interpretations of the projected test for Mandalay painting
- Categories:
 249 Views
249 ViewsA craniometry study was undertaken to obtain anthropometric measurements of three hundred and five (305) medical staff within Trinidad & Tobago which is a twin island republic situated in the Caribbean. A non-contact measurement method was used involving 3D scanning equipment to record the geometry of each subject’s head as a digital file. The digital files were then processed using CAD software to obtain measurements for twenty-two (22) facial points of interest. In addition, the gender of each staff member was recorded.
- Categories:
 254 Views
254 Views
Drug development is a process that is incredibly expensive and time-consuming. Computational drug repurposing can help to assign new indications for approved drugs. It is capable to reduce the cost of drug developments. Machine learning models have been introduced to repurpose drugs long before. Recent studies formulate computational drug repurposing problem as a latent link prediction task on a heterogenous network. A number of computational methods have been developed based on graph neural networks.
- Categories:
 104 Views
104 ViewsThe emergence of SARS-CoV-2 lineages derived from Omicron, including BA.2.86 (nicknamed “Pirola”) and its relative, JN.1, has raised concerns about their potential impact on public and personal health due to numerous novel mutations. Despite this, predicting their implications based solely on mutation counts proves challenging. Empirical evidence of JN.1’s increased immune evasion capacity in relation to previous variants is mixed.
- Categories:
 128 Views
128 ViewsThis dataset contains RF (Radio Frequency) signals obtained from simulations, which model ultrasound propagation in cortical bone.
The simulations were designed to provide insights into the behaviour of ultrasound waves in cortical bone tissues, both in intact and pathological conditions. The dataset covers a wide range of parameters, including varying thickness (1-8 mm), porosity (1-20%), and frequency (1-8 MHz), allowing to explore the impact of these factors on ultrasound signal characteristics.
- Categories:
 393 Views
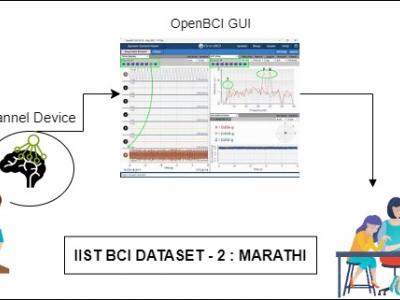
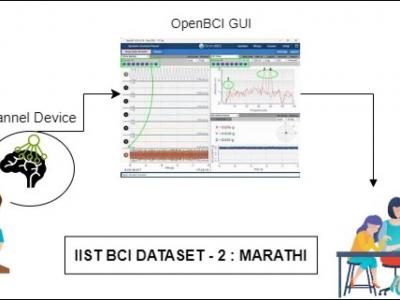
393 ViewsProblems of neurodegenerative disorder patients can be solved by developing Brain-Computer Interface (BCI) based solutions. This requires datasets relevant to the languages spoken by patients. For example, Marathi, a prominent language spoken by over 83 million people in India, lacks BCI datasets based on the language for research purposes. To tackle this gap, we have created a dataset comprising Electroencephalograph (EEG) signal samples of selected common Marathi words.
- Categories:
 526 Views
526 Views
Background: The endpoint of liposuction surgery in a current clinical dilemma. The liposuction volume (LV) should be individualized according to each patient's objective and subjective needs.
Objectives: To create a mathematical model to quantitatively predict the change ratio of thigh circumference to LV.
- Categories:
 83 Views
83 ViewsMS-BioGraphs are a family of sequence similarity graph datasets with up to 2.5 trillion edges. The graphs are weighted edges and presented in compressed WebGraph format. The dataset include symmetric and asymmetric graphs. The largest graph has been created by matching sequences in Metaclust dataset with 1.7 billion sequences. These real-world graph dataset are useful for measuring contributions in High-Performance Computing and High-Performance Graph Processing.
- Categories:
 765 Views
765 ViewsHealthcare 4.0 introduces the groundbreaking notion of the "Digital Twin (DT)," utilizing a digital model to encompass an individual's biological traits. This technology allows for the development of tailored treatment approaches, supports timely interventions, monitors respiratory issues, and offers decision-making assistance to healthcare professionals, ultimately advancing healthcare capabilities.
- Categories:
 428 Views
428 Views