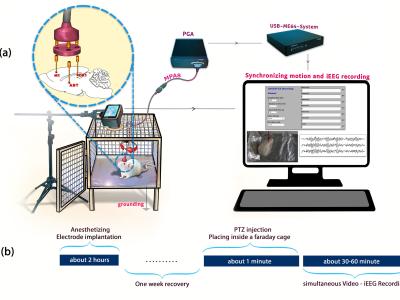
Fourteen participants completed this experiment. All participants were university students, physically healthy, with no history of neurological or psychiatric disorders, and were all right-handed. Written informed consent was obtained from each participant after a detailed explanation of the study protocol. All procedures were conducted in accordance with the Declaration of Helsinki. Due to issues encountered during the data collection process, the data from Participant S001 is deemed unreliable and has been excluded from further analysis.
- Categories: