Biomedical and Health Sciences

Subjects are categorized into three groups based on office blood pressure threshold: Normal (N), Prehypertension (P), and Stage 1 Hypertension (S). Each group contains 100 subjects, and all records have duration of at least 8 minutes. This study uses sliding window with length of 1 second and step size of 1 second to segment records. PPG, ECG and BP yield 167432 segments, respectively. MAP, DBP, and SBP are defined as average, minimum, and maximum of each BP segment, respectively. Max-Min normalization is applied to PPG and ECG segments.
- Categories:
 502 Views
502 Views
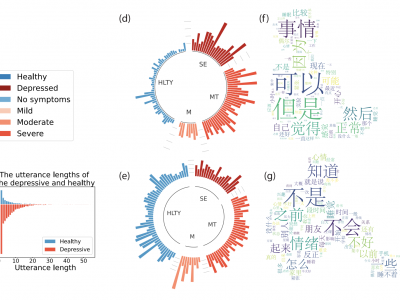
While functional near-infrared spectroscopy (fNIRS) had previously been suggested for major depressive disorder (MDD) diagnosis, the clinical application to predict antidepressant treatment response (ATR) is still unclear. To address this, the aim of the current study is to investigate MDD ATR using fNIRS and micro-ribonucleic acids (miRNAs). Our proposed algorithm includes a custom inter-subject variability reduction based on the Principal Component Analysis (PCA).
- Categories:
 177 Views
177 Views
The increased exposure of insects to radio frequency electromagnetic fields (RF-EMFs) may have an impact on their health. The RF-EMF absorbed power in certain insects is considerably higher in the range of 6-300 GHz, due to more comparable wavelengths to their size. Likewise, in this range, the near-field interactions between antennas' and certain insects can significantly affect antennas' performance.
- Categories:
 148 Views
148 Views
we searched the circRic database to gather data on the correlations between circRNAs and drug sensitivities, while the GDSC database contains information about drug sensitivities. The initial data set covers 80,076 associations related to 404 circRNAs and 250 drugs. The Wilcoxon test was used to determine the significant circRNA-drug sensitivity correlations. The procedure was contingent upon the false discovery rate (FDR) being less than 0.05.
- Categories:
 71 Views
71 Views
Background
Effective retraining of foot elevation and forward propulsion is essential in stroke survivors’ gait rehabilitation. However, home-based training often lacks valuable feedback. eHealth solutions based on inertial measurement units (IMUs) could offer real-time feedback on fundamental gait characteristics. This study aimed to investigate the effect of providing real-time feedback through an eHealth solution on foot strike angle (FSA) and forward propulsion in people with stroke.
- Categories:
 27 Views
27 ViewsThis dataset consists of 462 field of views of Giemsa(dye)-stained and field(dye)-stained thin blood smear images acquired using an iPhone 10 mobile phone with a 12MP camera. The phone was attached to an Olympus microscope with 1000× objective lens. Half of the acquired images are red blood cells with a normal morphology and the other half have a Rouleaux formation morphology.
- Categories:
 1014 Views
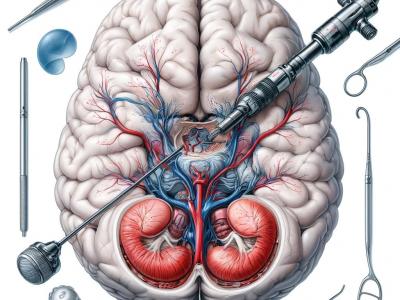
1014 ViewsThis dataset comprises 1718 annotated images extracted from 29 video clips recorded during Endoscopic Third Ventriculostomy (ETV) procedures, each captured at a frame rate of 25 FPS. Out of these images, 1645 are allocated for the training set, while the remainder is designated for the testing set. The images contain a total of 4013 anatomical or intracranial structures, annotated with bounding boxes and class names for each structure. Additionally, there are at least three language descriptions of varying technicality levels provided for each structure.
- Categories:
 418 Views
418 Views
We developed, implemented, and assessed the performance of two forms of plug-in type repetitive controllers (RC) for enhancing the transparency of a lower extremity exoskeleton that operates to support walking function. One controller is a first order RC (SING) consisting of a single period matched to the self-selected cadence of the participant. The second is a novel 'parallel' RC (PARA) which consists of a library of integrated RCs with varying periods, intended to accommodate a wider range of gait cycle times.
- Categories:
 86 Views
86 ViewsEarly detection of kidney illness can be achieved by training machine learning algorithms to discover patterns in patient data, such as imaging, test results, and medical history. This will enable rapid diagnosis and start of treatment regimens, which can improve patient outcomes. With 98.97% accuracy in CKD detection, the suggested TrioNet with KNN imputer and SMOTE fared better than other models. This comprehensive research highlights the model's potential as a useful tool in the diagnosis of chronic kidney disease (CKD) and highlights its capabilities.
- Categories:
 1198 Views
1198 Views