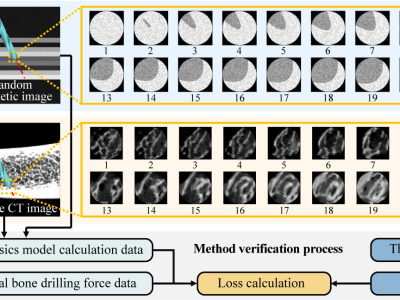
All the healthcare facilites in this dataset were collected from the MOH 2018 list of Uganda healthcare facilites (https://library.health.go.ug/sites/default/files/resources/National%20Health%20Facility%20MasterLlist%202017.pdf) Additional features were scraped using the Google Maps API and additionally from some of the websites of the healthcare facilities themselves.
- Categories: