Medical Imaging
This robust dataset is extracted from the International Skin Imaging Collaboration (ISIC). Similar datasets are used for the annual ISIC Challenge, presenting an opportunity for the computer science community to produce algorithms that can outperform professional dermatology. The submitted dataset contains approximately 1,000 images of malignant melanomas, as well as approximately 1,000 images of benign melanomas.
- Categories:
 3493 Views
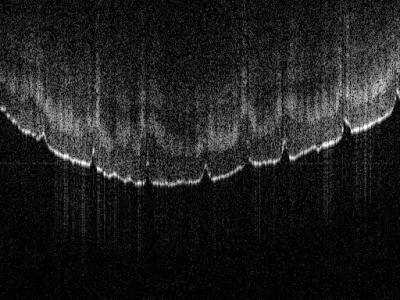
3493 ViewsThe dataset contains ungated free-breathing cardiac MRI scans with different image resolutions and slice thicknesses. Specifically, two low-resolution scans (2.27mm x 2.26mm resolution at 264 x 186 acquisition matrix size) are available with 10mm slice thickness and isotropic slice thickness, respectively, and a high-resolution scan (1.25mm x 1.26mm resolution at 480 x 334 acquisition matrix size) with 5mm slice thickness.
- Categories:
 1469 Views
1469 Views
Normal
0
7.8 磅
0
2
false
false
false
EN-US
ZH-CN
X-NONE
- Categories:
 54 Views
54 ViewsObjective: Stereoelectroencephalography (SEEG) is an established invasive diagnostic technique for use in patients with drug-resistant focal epilepsy evaluated before resective epilepsy surgery. The factors that influence the accuracy of electrode implantation are not fully understood. Adequate accuracy prevents the risk of major surgery complications. Precise knowledge of the anatomical positions of individual electrode contacts is crucial for the interpretation of SEEG recordings and subsequent surgery.
- Categories:
 384 Views
384 ViewsThree exemplary knee bone models derived from ultrasound imaging and their respective magnetic resonance imaging reference.
Apart from the ground truth, a partial scan as accessable by ultrasound imaging as well as full bone model computed by a statistical shape model is provided.
- Categories:
 543 Views
543 ViewsOvarian cancer is among the top health issues faced by women everywhere in the world . Ovarian tumours have a wide range of possible causes. Detecting and tracking down these cancers in their early stages is difficult which adds to the difficulty of treatment. In most cases, a woman finds out she has ovarian cancer after it has already spread. In addition, as technology in the field of artificial intelligence advances, detection can be done at an earlier level. Having this data will assist the gynaecologist in treating these tumours as soon as possible.
- Categories:
 3129 Views
3129 Views
This folder consists of codes, dataset, and models.
- Categories:
 29 Views
29 ViewsThis dataset consists of both non-retinal detachment and rhegmatogenous retinal detachment fundus images. The fundus images were collected from the four eye hospital in the country (namely India) such as Silchar medical college and hospital (Assam), Aravind eye hospital (Tamil Nadu), LV prasad eye hospital (Hyderabad), and Medanta- The medicity (Gurugram). A total of 1693 images have been collected from these hospitals of which 1017 fundus images belonged to retinal detachments and the rest 676 were non-retinal detachments.
- Categories:
 1755 Views
1755 Views