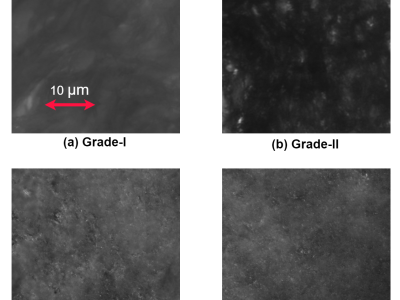
Early-stage cervical cancer is characterized by morphological changes in cells, which are effectively identified through biopsy techniques with high diagnostic accuracy. However, biopsies can be expensive and occasionally painful, with diagnosis reports often taking days or weeks to generate. These delays and costs create significant barriers for underprivileged women with limited access to timely and affordable healthcare.
- Categories: