Medical Imaging
- Categories:
 809 Views
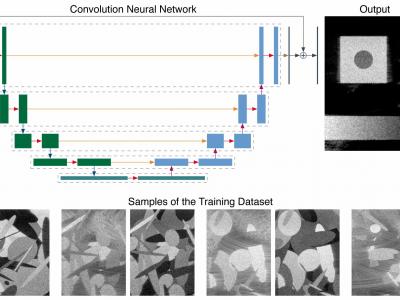
809 ViewsThis repository contains the data related to the paper “CNN-Based Image Reconstruction Method for Ultrafast Ultrasound Imaging” (10.1109/TUFFC.2021.3131383). It contains multiple datasets used for training and testing, as well as the trained models and results (predictions and metrics). In particular, it contains a large-scale simulated training dataset composed of 31000 images for the three different imaging configuration considered (i.e., low quality, high quality, and ultrahigh quality).
- Categories:
 3407 Views
3407 Views
Re-curated Breast Imaging Subset DDSM Dataset (RBIS-DDSM) is a curated version of 849 images from the CBIS-DDSM dataset available online with a permissive copyright license (CC-BY-SA 3.0). The CBIS-DDSM dataset is an improved version of the DDSM dataset. The authors of the CBIS-DDSM dataset attempted to improve the ground truth by applying simple image processing based methods to enhance the edges without any manual intervention from medical experts in order to segment and annotate masses. However, these annotations (segmentation maps) are inaccurate in most of the images.
- Categories:
 967 Views
967 Views
The Femoropopliteal Artery Stent Dataset is captured by Siemens digital radiography fluoroscopy system Luminos dRF Max. The dataset is from the First Affiliated Hospital of Fujian Medical University in Fuzhou City (Fujian Province, China). The X-ray fluoroscopic image data set of the femoropopliteal artery stent was obtained from different positions of the patient.
The provided dataset file contains the following directory structure when unzipped:
- Categories:
 322 Views
322 Views
Reconstructed 2D Shepp-Logan phantom and brain MRA projected image using different flat portions in pulsed excitation and magnetic nanoparticle sizes.
- Categories:
 51 Views
51 ViewsThe Open Big Healthy Brains (OpenBHB) dataset is a large (N>5000) multi-site 3D brain MRI dataset gathering 10 public datasets (IXI, ABIDE 1, ABIDE 2, CoRR, GSP, Localizer, MPI-Leipzig, NAR, NPC, RBP) of T1 images acquired across 93 different centers, spread worldwide (North America, Europe and China). Only healthy controls have been included in OpenBHB with age ranging from 6 to 88 years old, balanced between males and females.
- Categories:
 9912 Views
9912 Views
An enhanced dataset, VQA-RADPh, based on the VQA-RAD dataset.
- Categories:
 161 Views
161 Views