CNN-Based Image Reconstruction Method for Ultrafast Ultrasound Imaging: Data
- Citation Author(s):
-
Jean-Philippe Thiran (École polytechnique fédérale de Lausanne (EPFL))
- Submitted by:
- Dimitris Perdios
- Last updated:
- DOI:
- 10.21227/vn0e-cw64
- Data Format:
 3506 views
3506 views
- Categories:
- Keywords:
Abstract
This repository contains the data related to the paper “CNN-Based Image Reconstruction Method for Ultrafast Ultrasound Imaging” (10.1109/TUFFC.2021.3131383). It contains multiple datasets used for training and testing, as well as the trained models and results (predictions and metrics). In particular, it contains a large-scale simulated training dataset composed of 31000 images for the three different imaging configuration considered (i.e., low quality, high quality, and ultrahigh quality). It also contains images of a dedicated numerical test phantom (300 realizations) associated to ultrasound-specific image metrics, 300 additional (test) samples simulated identically to the training dataset, an in vivo test dataset acquired on the carotid of a volunteer (60 frames, longitudinal view), and an in vitro test dataset acquired on a CIRS model 054GS.
The accepted version of this paper is also available on arXiv: arXiv:2008.12750.
The corresponding code is available online at https://github.com/dperdios/dui-ultrafast.
Paper Abstract
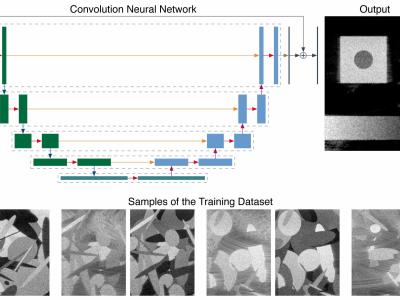
Ultrafast ultrasound (US) revolutionized biomedical imaging with its capability of acquiring full-view frames at over 1 kHz, unlocking breakthrough modalities such as shear-wave elastography and functional US neuroimaging. Yet, it suffers from strong diffraction artifacts, mainly caused by grating lobes, side lobes, or edge waves. Multiple acquisitions are typically required to obtain a sufficient image quality, at the cost of a reduced frame rate. To answer the increasing demand for high-quality imaging from single unfocused acquisitions, we propose a two-step convolutional neural network (CNN)-based image reconstruction method, compatible with real-time imaging. A low-quality estimate is obtained by means of a backprojection-based operation, akin to conventional delay-and-sum beamforming, from which a high-quality image is restored using a residual CNN with multiscale and multichannel filtering properties, trained specifically to remove the diffraction artifacts inherent to ultrafast US imaging. To account for both the high dynamic range and the oscillating properties of radio frequency US images, we introduce the mean signed logarithmic absolute error (MSLAE) as a training loss function. Experiments were conducted with a linear transducer array, in single plane-wave (PW) imaging. Trainings were performed on a simulated dataset, crafted to contain a wide diversity of structures and echogenicities. Extensive numerical evaluations demonstrate that the proposed approach can reconstruct images from single PWs with a quality similar to that of gold-standard synthetic aperture imaging, on a dynamic range in excess of 60 dB. In vitro and in vivo experiments show that trainings carried out on simulated data perform well in experimental settings.
License
Data released under the terms of the Creative Commons Attribution 4.0 International (CC BY 4.0).
If you are using this data and/or code, please cite the corresponding paper.
Acknowledgments
This work was supported in part by the Swiss National Science Foundation under Grant 205320_175974 and Grant 206021_170758.
Instructions:
The detailed description of the data available in this repository can be found online at https://github.com/dperdios/dui-ultrafast/#data.
Dataset Files
- 20200304-ge9ld-random-phantom.hdf5 (Size: 36.25 KB)
- 20200304-ge9ld-random-phantom.images.hq.hdf5 (Size: 220.25 GB)
- 20200304-ge9ld-random-phantom.images.lq.hdf5 (Size: 220.25 GB)
- 20200304-ge9ld-random-phantom.images.uq.hdf5 (Size: 220.25 GB)
- 20200304-ge9ld-numerical-test-phantom.hdf5 (Size: 36.25 KB)
- 20200304-ge9ld-numerical-test-phantom.images.hq.hdf5 (Size: 2.13 GB)
- 20200304-ge9ld-numerical-test-phantom.images.lq.hdf5 (Size: 2.13 GB)
- 20200304-ge9ld-numerical-test-phantom.images.uq.hdf5 (Size: 2.13 GB)
- 20200304-ge9ld-numerical-test-phantom-predictions.hdf5 (Size: 8.53 GB)
- 20200304-ge9ld-random-phantom-test-set.hdf5 (Size: 38.32 KB)
- 20200304-ge9ld-random-phantom-test-set.images.hq.hdf5 (Size: 2.13 GB)
- 20200304-ge9ld-random-phantom-test-set.images.lq.hdf5 (Size: 2.13 GB)
- 20200304-ge9ld-random-phantom-test-set.images.uq.hdf5 (Size: 2.13 GB)
- 20200304-ge9ld-random-phantom-test-set.inclusions.amp.hdf5 (Size: 254.63 KB)
- 20200304-ge9ld-random-phantom-test-set.inclusions.ang.hdf5 (Size: 254.63 KB)
- 20200304-ge9ld-random-phantom-test-set.inclusions.pos.hdf5 (Size: 726.94 KB)
- 20200304-ge9ld-random-phantom-test-set.inclusions.semiaxes.hdf5 (Size: 492.56 KB)
- 20200304-ge9ld-random-phantom-test-set-predictions.hdf5 (Size: 5.33 GB)
- 20200527-ge9ld-experimental-test-set-carotid-long.hdf5 (Size: 26.23 KB)
- 20200527-ge9ld-experimental-test-set-carotid-long.images.hq.hdf5 (Size: 436.54 MB)
- 20200527-ge9ld-experimental-test-set-carotid-long.images.lq.hdf5 (Size: 436.54 MB)
- 20200527-ge9ld-experimental-test-set-carotid-long-predictions.hdf5 (Size: 436.54 MB)
- 20200527-ge9ld-experimental-test-set-cirs054gs-hypo2.hdf5 (Size: 26.23 KB)
- 20200527-ge9ld-experimental-test-set-cirs054gs-hypo2.images.hq.hdf5 (Size: 7.29 MB)
- 20200527-ge9ld-experimental-test-set-cirs054gs-hypo2.images.lq.hdf5 (Size: 7.29 MB)
- 20200527-ge9ld-experimental-test-set-cirs054gs-hypo2-predictions.hdf5 (Size: 7.29 MB)
- metrics.tar.gz (Size: 51.18 MB)
- trained-models.tar.gz (Size: 1.53 GB)







good job!!