Brain
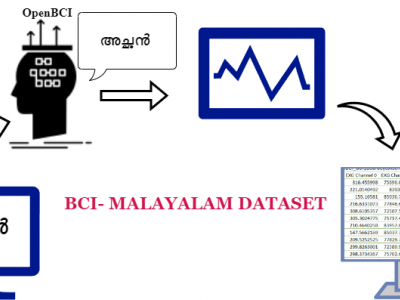
This paper presents a dataset of brain Electroencephalogram (EEG) signals created when Malayalam vowels and consonants are spoken. The dataset was created by capturing EEG signals utilizing the OpenBCI Cyton device while a volunteer spoke Malayalam vowels and consonants. It includes recordings obtained from both sub-vocal and vocal. The creation of this dataset aims to support individuals who speak Malayalam and suffer from neurodegenerative diseases.
- Categories:
 2591 Views
2591 Views
This dataset acompanies our article titled "Insights into traditional Large Deformation Diffeomorphic Metric Mapping and unsupervised deep-learning for diffeomorphic registration and their evaluation", Computers in Biology and Medicine, 2024. This paper explores the connections between traditional Large Deformation Diffeomorphic Metric Mapping methods and unsupervised deep-learning approaches for non-rigid registration, particularly emphasizing diffeomorphic registration.
- Categories:
 271 Views
271 Views
Traditional authentication models are vulnerable to security breaches when personal data is exposed. This study introduces novel hybrid visual stimuli protocols integrating event-related potentials (ERP) and steady-state visually evoked potentials (SSVEP) to develop an authentication system that enhances both performance and personalization in neural interfaces. Our model utilizes distinctive neural patterns elicited by a range of visual stimuli based on 4-digit numbers, such as familiar numbers (personal birthdates, excluding targets), standard targets, and non-targets.
- Categories:
 32 Views
32 ViewsTo address the challenges faced by patients with neurodegenerative disorders, Brain-Computer Interface (BCI) solutions are being developed. However, many current datasets lack inclusion of languages spoken by patients, such as Telugu, which is spoken by over 90 million people in India. To bridge this gap, we have created a dataset comprising Electroencephalograph (EEG) signal samples of commonly used Telugu words. Using the Open-BCI Cyton device, EEG samples were captured from volunteers as they pronounced these words.
- Categories:
 466 Views
466 ViewsThis paper introduces a dataset capturing brain signals generated by the recognition of 100 Malayalam words, accompanied by their English translations. The dataset encompasses recordings acquired from both vocal and sub-vocal modalities for the Malayalam vocabulary. For the English equivalents, solely vocal signals were collected. This dataset is created to help Malayalam speaking patients with neuro-degenerative diseases.
- Categories:
 2859 Views
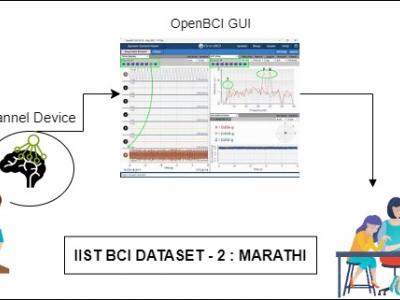
2859 ViewsProblems of neurodegenerative disorder patients can be solved by developing Brain-Computer Interface (BCI) based solutions. This requires datasets relevant to the languages spoken by patients. For example, Marathi, a prominent language spoken by over 83 million people in India, lacks BCI datasets based on the language for research purposes. To tackle this gap, we have created a dataset comprising Electroencephalograph (EEG) signal samples of selected common Marathi words.
- Categories:
 527 Views
527 ViewsThis dataset consists of electroencephalography (EEG) data from 6 participants aged between 23 and 28 years, with a mean age of 25 years. The dataset is the motor imagery EEG signals of six different rehabilitation training movements in the upper limbs. We recruited six participants aged between 23 and 28 years, with a mean age of 25 years. Three of the participants are male. Subjects sat in a comfortable chair, facing the computer monitor that displayed the trial-based paradigm and their right arm was naturally placed on the table to avoid muscle fatigue.
- Categories:
 658 Views
658 Views
Faces and bodies provide critical cues for social interaction and communication. Their structural encoding depends on configural processing, as suggested by the detrimental effect of stimulus inversion for both faces (i.e., face inversion effect - FIE) and bodies (body inversion effect - BIE). An occipito-temporal negative event-related potential (ERP) component peaking around 170 ms after stimulus onset (N170) is consistently elicited by human faces and bodies and it is affected by the inversion of these stimuli.
- Categories:
 319 Views
319 ViewsIn today’s context, it is essential to develop technologies to help older patients with neurocognitive disorders communicate better with their caregivers. Research in Brain Computer Interface, especially in thought-to-text translation has been carried out in several languages like Chinese, Japanese and others. However, research of this nature has been hindered in India due to scarcity of datasets in vernacular languages, including Malayalam. Malayalam is a South Indian language, spoken primarily in the state of Kerala by bout 34 million people.
- Categories:
 930 Views
930 Views