Datasets
Open Access
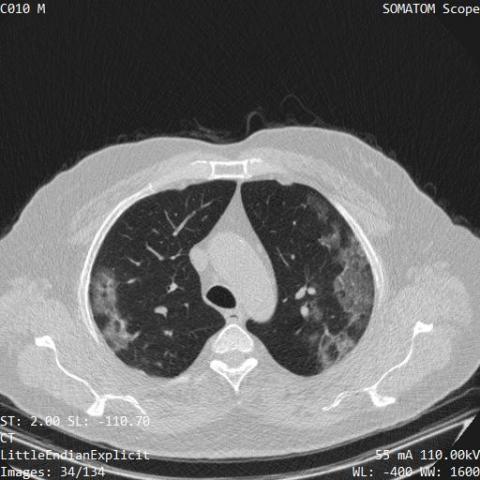
COVID-19 Low-Dose and Ultra-Low-Dose CT Scans
- Citation Author(s):
- Submitted by:
- Shahin Heidarian
- Last updated:
- Tue, 06/01/2021 - 10:21
- DOI:
- 10.21227/sed8-6r15
- Links:
- License:
 3209 Views
3209 Views- Categories:
- Keywords:
Abstract
Reverse transcription-polymerase chain reaction (RT-PCR) is currently the gold standard in COVID-19 diagnosis. It can, however, take days to provide the diagnosis, and false negative rate is relatively high. Imaging, in particular chest computed tomography (CT), can assist with diagnosis and assessment of this disease. Nevertheless, it is shown that standard dose CT scan gives significant radiation burden to patients, especially those in need of multiple scans. Therefore, we collect low-dose and ultra-low-dose (LDCT and ULDCT) scan protocols that reduce the radiation exposure close to that of a single X-Ray, while maintaining an acceptable resolution for diagnosis purposes. This dataset contains scans of 104 COVID-19 positive cases, and 56 normal cases, collected in Babak Imaging Center, Tehran, Iran.
36.5% of the COVID-19 cases are confirmed with positive RT-PCR. Diagnosis for the rest of the cases is obtained by consensus between three experienced radiologists with 95.6% agreement. Slices level labels are also provided to specify the slices demonstrating infection in the CT scans.
This dataset also includes an extra set of 100 positive COVID-19 patients, confirmed with positive RT-PCR with or without related COVID-19 lung manifestations.
The volumetric LDCT and ULDCT scans are obtained from a SIEMENS SOMATOM Scope scanner. All scans are in the axial view and reconstructed into 512,512 images using the Filtered Back Projection method. The radiation dose in standard chest CT scans is estimated at 7 mSv, which is reduced to 1–1.5 mSv in LDCT scans and as low as 0.3 mSv in the ULDCT ones. For patients with >60 kg body weight LDCT images are acquired using the mAs value of 20, kVp of 110 v, and the slice thickness of 2 mm, whereas for patients with the body weight of less than 60 kg the ULDCT images are obtained with 15 mAs. All cases are accompanied by demographic and clinical data, i.e., sex, age, weight, and presence or absence of 5 symptoms of cough, fever, dyspnea, chest pain, and fatigue.
“Dataset-S1” contains two folders for COVID-19 and Normal DICOM images, named as “COVID-S1” and “Normal-S1”, respectively. Within the same folder, three CSV files are available. The first one, named as “Radiologist-S1.csv”, contains labels assigned to the corresponding cases by three experienced radiologists. The second CSV file, “Clinical-S1.csv”, includes the clinical information as well as the result of the RT-PCR test, if available. The third file is named “LDCT-SL-Labels-S1.csv” and contains the slice-level labels related to COVID-19 cases. In other words, slices demonstrating infection are specified in this file.
Each row in this CSV file corresponds to a specific case, and each column represents the slice number in the volumetric CT scan. Label 1 indicates a slice with the evidence of infection, while 0 is assigned to slices with no evidence of infection.
Note that slices in each case should be sorted based on the “Slice-Location” value to match with the provided labels in the CSV file. The Slice Location values are stored in DICOM files and accessible from the following DICOM tag:
(0020,1041) – DS – Slice Location
“Dataset-S2” contains 100 COVID-19 positive cases, confirmed with RT-PCR test. 68 cases have related imaging findings, whereas 32 do not reveal signs of infection. These two groups are placed in two folders of “PCP-Lung-Positive “and “PCP-Lung-Negative”. “Dataset-S2” also includes a CSV file, namely “Clinical-S2.csv” presenting the clinical information.
Dataset Files
COVID-LDCT.zip (10.63 GB)
Open Access dataset files are accessible to all logged in users. Don't have a login? Create a free IEEE account. IEEE Membership is not required.