Datasets
Standard Dataset
DATASET for "Brain Injury Localization and stroke classification in Electromagnetic Imaging Using Graph Approaches"
- Citation Author(s):
- Submitted by:
- Guohun Zhu
- Last updated:
- Sat, 11/19/2022 - 01:20
- DOI:
- 10.21227/es43-bf10
- Data Format:
- License:
 4828 Views
4828 Views- Categories:
- Keywords:
Abstract
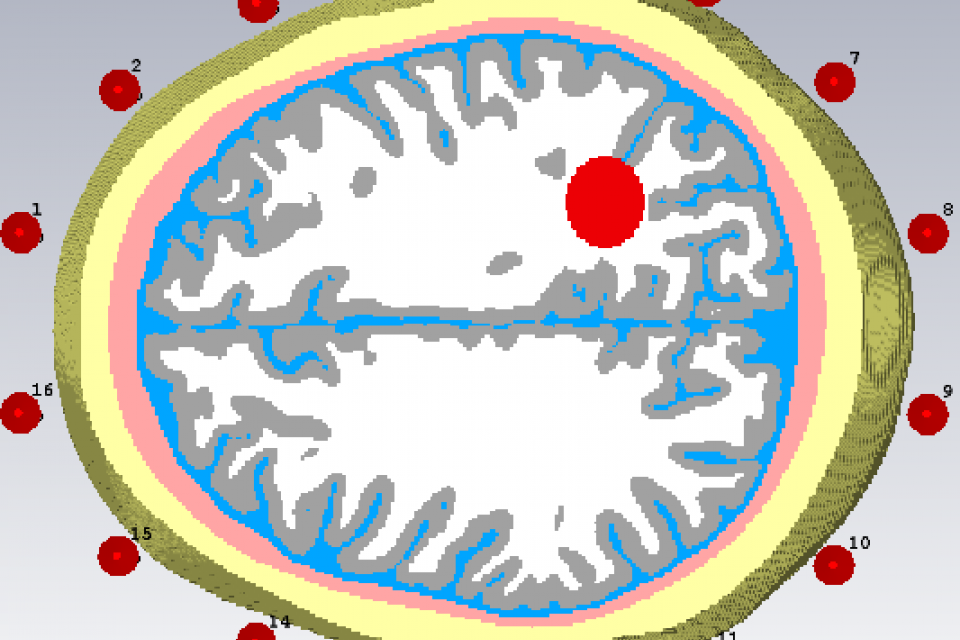
The proposed signals are used for electromagnetic-based stroke classification. Six realistic head phantom computed from MRI scans, is surrounded by an antenna array of 16 dipole antennas distributed uniformly around the head. These antennas are deployed in a fixed circular array around the head, at a distance of approximately 2-3 mm from the head. A Gaussian pulse covering the bandwidth from 0:7 to 2 GHz is emitted from each of the antennas, sequentially, while all of the antennas capture the scattered signals. Since 16 antennas were used, there are a total of 256 channel signals (i.e. 16x16) as input signals.
This data is used to test classficiation for stoke types. Brain stroke can be classified into two categories; intra-cranial haemorrhage (ICH) and ischemic stroke (IS). ICH is caused by the rupture of a blood vessel, whereas IS results from a clot which restricts the blood flow to brain tissues. Although both types of strokes need emergency treatment, the treatment of a stroke depends upon the type of stroke (e.g. ischemic or haemorrhagic). It is not only important that the type of stroke is diagnosed quickly to reduce the brain damage, but also because a treatment suitable for one type of stroke may be harmful when treating a different type. Thus, identifying the stroke types are an important issue in a microwave system.
How to show the touchstone files?
SNP files can be open by many tools. Here I recommend two methods.
(1) ReadSNP: it is coded by Java Language. You can use it to visualize 16 magnitude S-parameter scatter signals from an antenna. This tool can be downloaded from the following link.
http://uadi.project.uq.edu.au/UADI/stroke/ReadSNP.zip
(2)Scikit-rf: It is a RF/Microwave engineering package implemented in the Python programming language. The details can be found in the following link.
https://scikit-rf.readthedocs.io/en/latest/tutorials/Introduction.html
If you want to understand the SNP file format in details, you can read the SnP(Touchstone) File Format from the keysight website.
If you need the further details, please contact the author: g.zhu at uq.edu.au
More from this Author