TXT
In order to accelerate the CU partitioning in VVC-SCC, we have established a specialized dataset for screen content. Firstly, we collect a wide variety of screen content videos and images in YUV format. Secondly, we encode these sequences under four different Quantization Parameters (QPs) to obtain comprehensive information about CU partitioning in VVC-SCC. This includes the length, width, position, mode labels, and RDcost values corresponding to each mode. This dataset will provide valuable data support for optimizing the CU partitioning process in screen content coding.
- Categories:
 74 Views
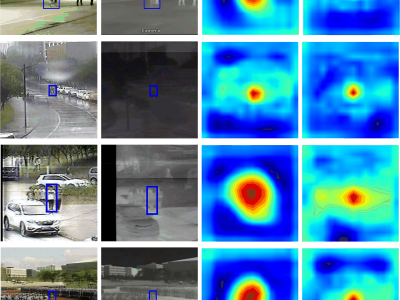
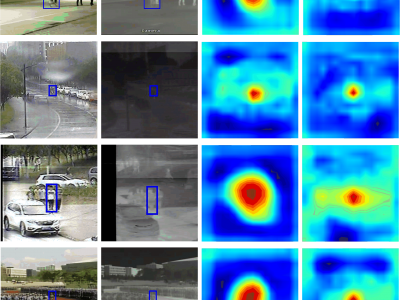
74 ViewsWe are pleased to submit our manuscript entitled "Hetero-modal Template Guide Search Region for RGBT Tracking" for consideration for publication in IEEE Transactions on Consumer Electronics. This work presents a novel framework for robust RGB-Thermal (RGBT) object tracking, addressing critical challenges in consumer electronics applications such as smart security systems, autonomous navigation, and augmented reality devices.
- Categories:
 173 Views
173 ViewsWe are pleased to submit our manuscript entitled "Hetero-modal Template Guide Search Region for RGBT Tracking" for consideration for publication in IEEE Transactions on Consumer Electronics. This work presents a novel framework for robust RGB-Thermal (RGBT) object tracking, addressing critical challenges in consumer electronics applications such as smart security systems, autonomous navigation, and augmented reality devices.
- Categories:
 138 Views
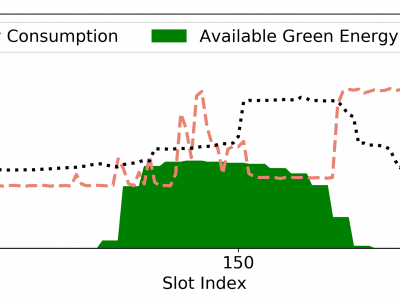
138 ViewsData centers are increasingly adopting renewable energy sources to mitigate environmental impact and reduce operational costs. However, effectively optimizing energy costs remains challenging due to unpredictable workloads and fluctuating renewable energy availability. This paper introduces LOECM, a Lyapunov-driven online scheduling algorithm designed to minimize energy cost without relying on future information.
- Categories:
 56 Views
56 Views
This dataset was synthesized through a large model. We deployed two large models, one playing the role of an elderly patient and the other playing the role of a nurse. The scenario we define is as follows: the patient lies on a deformed bed, conveying clear or unclear needs to the nurse. The nurse analyzes the patient's true needs based on their language, provides a humanized and personalized response to the patient, and issues control commands to the deformed bed. The dataset consists of two parts: patient statements - answers and patient statements - control commands.
- Categories:
 24 Views
24 ViewsThis dataset comprises images of parts from real industrial scenarios and virtual reality environments. Real images are sourced from actual industrial settings, ensuring both authenticity and diversity, while virtual reality images, which make up approximately 11% of the dataset, are captured through precise 3D modeling. Approximately 30% of the part information was manually authored by industry experts, while the remaining 70% was generated by multimodal large models such as Wenxin Yiyan and GPT-4.
- Categories:
 197 Views
197 Views
Numerous studies have demonstrated that microbes play a vital role in human health, making the identification of potential microbe-drug associations critical for drug discovery and clinical treatment. In this manuscript, we proposed a novel prediction model named GTDEKAN by integrating an aware Transformer network with a Dual Cross-Attention (DCA) module (including a Channel Cross-Attention and a Spatial Cross-Attention) and an Enhanced Kolmogorov-Arnold Network (EKAN) to infer potential microbe-drug associations.
- Categories:
 59 Views
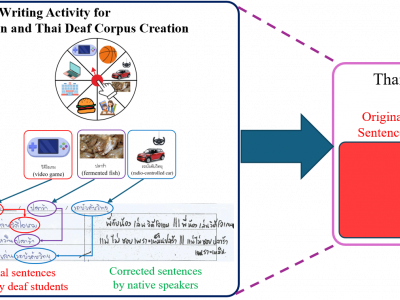
59 ViewsThe Thai Deaf Corpus (TDC) is constructed from a writing activity where deaf students randomly select picture words using the image picker wheel, then write sentences corresponding to these words on the writing sheet. The sentences are transcribed and corrected manually to create the TDC.
- Categories:
 254 Views
254 ViewsA dataset incldes 841 nodes. This dataset includes 841 nodes in a mobile social network, used to simulate the process of users being interconnected and influencing each other within the mobile social network. Each row of data consists of two numbers, representing the current location of the node. In the process of information dissemination, each user is a node, influenced by their neighboring nodes and also influencing those neighbors in return.
- Categories:
 370 Views
370 Views
This dataset comprises over 38,000 seed inputs generated from a range of Large Language Models (LLMs), including ChatGPT-3.5, ChatGPT-4, Claude-Opus, Claude-Instant, and Gemini Pro 1.0, specifically designed for the application in fuzzing Python functions. These seeds were produced as part of a study evaluating the utility of LLMs in automating the creation of effective fuzzing inputs, a method crucial for uncovering software defects in the Python programming environment where traditional methods show limitations.
- Categories:
 232 Views
232 Views