A Molecular Communication Testbed Based on Proton Pumping Bacteria
- Citation Author(s):
-
Laura GrebensteinJens KirchnerArman AhmadzadehVahid JamaliGeorg FischerRobert WeigelAndreas BurkovskiRobert Schober
- Submitted by:
- Wayan Wicke
- Last updated:
- DOI:
- 10.21227/3zj6-pm05
- Data Format:
- Research Article Link:
- Links:
 634 views
634 views
- Categories:
- Keywords:
Abstract
This data set is from a recent biological molecular communication (MC) testbed and provides a set of experimental measurement data.
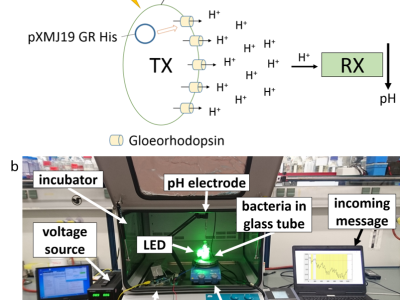
In particular, the MC testbed is realized using Escherichia coli (E. coli) bacteria that express the light-driven proton pump gloeorhodopsin (GR) from Gloeobacter violaceus (G. violaceus).
Upon an external light stimulus, these bacteria (as the transmitter) export protons (as the signaling molecules) into the channel.
This changes the pH level of the environment, which can be detected with a pH sensor (as the receiver).
The data set is expected to facilitate and promote the experimental evaluation of communication schemes developed by the MC community.
Instructions:
Package:
- Comprehensive description of the data set and experimental procedure: supplement.pdf
- Publication: A Molecular Communication Testbed Based on Proton Pumping Bacteria: Methods and Data, IEEE Transactions on Molecular, Biological, and Multi-Scale Communications
- Signals for 18 experiments of types "Dark Adaption", "Control Response", "Long On-Off Response", and "Bit Response": SignalData.zip
- Signals for 12 noise measurements: NoiseData.zip
- Matlab script for visualization: example_DisplayData.m
Citing:
- Reference to the experimental system: Biological Optical-to-Chemical Signal Conversion Interface: A Small-Scale Modulator for Molecular Communications, https://ieeexplore.ieee.org/document/8467351
- Reference to this data set: A Molecular Communication Testbed Based on Proton Pumping Bacteria: Methods and Data, IEEE Transactions on Molecular, Biological, and Multi-Scale Communications