Full Factorial Design of Experiment dataset of Silk Fibroin alkaly degumming
- Citation Author(s):
- Submitted by:
- Alessio Bucciarelli
- Last updated:
- DOI:
- 10.21227/emar-9460
- Data Format:
- Links:
 1284 views
1284 views
- Categories:
- Keywords:
Abstract
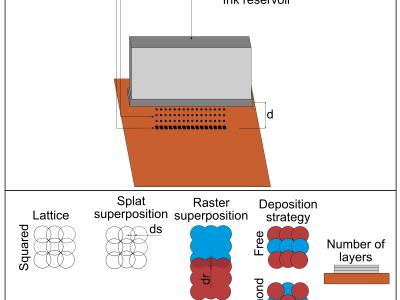
Silk fibroin is the structural fiber of the silk filament and it is usually separated from the external fibroin by a chemical process called degumming. This process consists in an alkali bath in which the silk cocoons are boiled for a determined time. It is also known that the degumming process impacts the property of the outcoming silk fibroin fibers. In this work we present the dataset obtained from a Design of Experiment (DoE) screening made on the alkali degumming and wich results were publisced ACS biomaterials science and Engineering (https://doi.org/10.1021/acsbiomaterials.0c01657). Four process factors were considered: the number of degumming baths, the process time, the process temperature and the salt concentration. The properties of the silk fibroin fibers were studied. In particular the molecular weight was obtained by gel permeation chromatography (GPC), the mechanical data by tensile test, the secondary structure by Fourier infrared spectroscopy (FTIR).
Instructions:
The data contained in the first sheet of the dataset is in tidy format (each row correspond to an observation) and can be directly imported in R and elaborated with the package Tidyverse. It should be noticed that the row with the standard order 49 correspond to the reference degumming while the row 50 correspond to the test made on the bare silk fiber (not degummed). In this last case neither the mass loss nor the secondary structures were determined. In fact, being not degummed the sericine was surrounding the fiber so the examination of the secondary structure could not be done. The first two column of the dataset represent the Standard order (the standard order in which the Design of Experiment data are elaborated) and the Run order (the randomized order in whcih the trials were performed). The next four columns are the Studied factors while the rest of the dataset reports the process yields (in this case, the properties of the outcoming silk fibers).
The second sheet contains the information of the molecular weight of the tested samples. In this case only one sample for each triplicate was tested. Both the standard order and the run order referred to the same samples of the first sheet.
In the Raw data.zip file the raw mechanical curves are reported in OriginLab format divided in datasheets numbered as the sample from 1 to 48 with the additions of a datasheet for the reference curves obtained form the Rokwood protocol and the curves form the raw cocoons.
In the same archive a file with the GPC curves and their elaborations for the tested samples are reported.