A Large-Scale Fully Annotated Foldscope Microscopy Image Dataset for Deep Learning Framework

- Citation Author(s):
-
Sumona BiswasShovan Barma
- Submitted by:
- sumona biswas
- Last updated:
- DOI:
- 10.21227/krmf-s880
- Data Format:
- Links:
 258 views
258 views
- Categories:
- Keywords:
Abstract
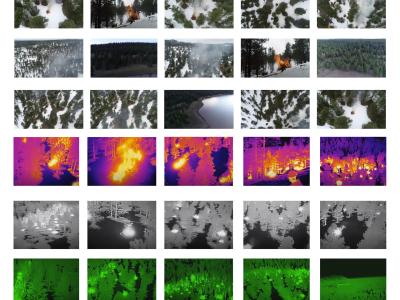
This work presents a large-scale three-fold annotated, low-cost microscopy image dataset of potato tubers for plant cell analysis in deep learning (DL) framework which has huge potential in the advancement of plant cell biology research. Indeed, low-cost microscopes coupled with new-generation smartphones could open new aspects in DL-based microscopy image analysis, which offers several benefits including portability, ease of use, and maintenance. However, its successful implications demand properly annotated large number of diverse microscopy images, which has not been addressed properly— that confines the advanced image processing based plant cell research. Therefore, in this work, a low-cost microscopy image database of potato tuber cells having a total 34,657 number of images, has been generated by Foldscope (costs around 1 USD) coupled with a smartphone. This dataset includes 13,369 unstained and 21,288 stained (safranin-o, toluidine blue-o, and lugol’s iodine) images with three-fold annotation based on weight, section areas, and tissue zones of the tubers.
Instructions:
In total, 34,657 of number of images including 13,369 unstained and 21,288 stained are generated by the Foldscope. The generated microscopy images are annotated by three-fold class labels based on the weight, section area, and tissue zone of the tubers. Firstly, the weight of the respective tubers is specified by L, M, and S for large, medium, and small respectively along with the sample number ranging from 1 to 5; for instance, L1 refers to the first sample of a large tuber. Secondly, three section areas, bud end, middle, and stem end are indicated by Z1, Z2, and Z3 respectively; and finally, the images captured from the inner and outer core are recorded as IC and OC respectively. Certainly, the image annotation information is inserted as an image file name by the abstractions corresponding to weight, section area, and tissue zone. Therefore, the file naming format of unstained images is as, weight with sample no._section area_tissue zone_section no._image no.; for example, a file name, L2_Z1_IC_Sec1_002.jpg indicates the second (002) unstained image of first section (Sec1) out of five, taken from the inner core (IC) of bud end (Z1) of the second sample of the large weight potato tuber (L2). Certainly, the dataset includes stained images which are generated by three different staining agents. Therefore, following the above file naming format, after section number information specific staining agent information has been embedded; for instance, M5_Z2_OC_Sec3_tolu_004.jpg, refers to an image stained by toluidine blue-o (tolu) and it is the fourth (004) stained image of the third section (Sec3) out of five, taken from the outer core (OC) of middle section (Z2) of the fifth sample of the medium weight potato tuber (M5). The dataset can be downloaded as a zip file. The dataset contains two folders named as “unstain” and “stain”. The “stain” and “unstain” folder contains raw microscopy images in JPG format. Three different stained images are in the respective folders named “safranin_o”, “toluidine_blue_o”, and “lugols_iodine”.







download dataset