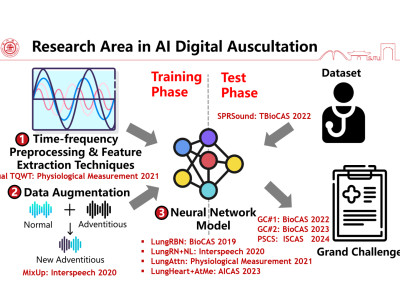
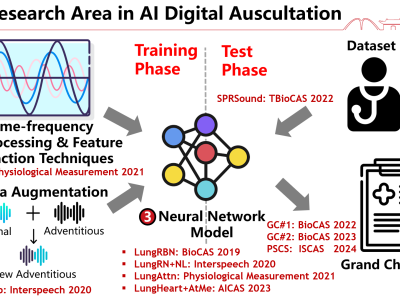
Respiratory Sound Track Grand Challenge 2023: Respiratory Sound Classification for SPRSound Dataset
- Submission Dates:
-
to
- Citation Author(s):
-
Qing Zhang (Shanghai Jiao Tong University)Jing ZhangJiajun YuanHuajie HuangYuhang ZhangChangyan ChenJilei LinBaoqin ZhangGaomei LvShuzhu LinNa WangXin LiuMingyu TangYahua WangLu LiuHui MaDan XieLihua WuHaibo YangShuhua YuanMengjun ChenBingxue ZhangHongyuan ZhouJian ZhaoYongfu LiYong YinLiebin ZhaoGuoxing WangYong Lian
- Submitted by:
- Yongfu Li
- Last updated:
- DOI:
- 10.21227/rqh3-h568
- Data Format:
- Links:
Abstract
Globally, respiratory diseases are the leading cause of death, making it essential to develop an automatic respiratory sounds software to speed up diagnosis and reduce physician workload. A recent line of attempts have been proposed to predict accurately, but they have yet been able to provide a satisfactory generalization performance. In this contest, we invited the community to develop more accurate and generalized respiratory sound algorithms. A starter code is provided to standardize the submissions and lower the barrier. New testing set is prepared to evaluate the generalization performance of the submissions. Top 3 teams will present their work at IEEE BioCAS 2023 conference.
Instructions:
About
According to the World Health Organization, respiratory diseases, such as pneumonia, asthma, bronchitis, lung cancer, and chronic obstructive pulmonary disease (COPD), are one of the most common mortality factors in the world, causing the death of more than 3 million people each year worldwide. These respiratory diseases have a direct impact on people’s social, economic, and health life. Early diagnosis is the key factor for preventing the spread of respiratory diseases and limiting the adverse effects on people’s life.
Auscultation is a technique where specialists use a stethoscope to detect adventitious lung sounds and identify the possible lung diseases. It provides a simple, low-cost, and non-invasive method for respiratory disease diagnosis. However, the auscultation technique based on stethoscope suffers from two disadvantages. Firstly, auscultation requires expert physicians to analyze the lung sounds. In impoverished environments with a shortage of expert physicians, any sudden and massive infectious respiratory diseases outbreak (such as the pneumonia complication from the coronavirus can further exacerbate the spread of the diseases and the increase of death rate. Secondly, even if the patients are diagnosed by experienced physicians, there might be subjectivity in the interpretations of lung sounds, causing inter-listener variability. Therefore, it is necessary to develop an automatic respiratory detection method to reduce the workload for physicians and eliminate subjectivity during diagnosis.
The IEEE BioCAS 2023 grand challenge on respiratory sound classification invites participants to explore different feature extraction techniques, model classification to improve the current state-of-the-art works. This new dataset is collected from the Shanghai Children’s Medical Center (SCMC), targeting children ranging from 1 month to 18 years old, which presents unseen challenges as compared to the prior datasets.
Timeline
| Start of Registration | Wednesday, March 1,2023 |
| Start of Project Submission | Friday, May 19,2023 |
| End of Project Submission/Regular Paper Submission Deadline | Friday, June 9,2023 |
| Author Notification Date | Friday, August 11, 2023 |
| BioCAS 2023 Student Travel Grants Application Deadline | Friday, August 24, 2023 |
| Author Registration/Final Paper Submission Deadline | Friday, September 1, 2023 |
| Conference Registration | Friday, October 6, 2023 |
Participation
- The competition is open to individuals, colleges/universities, scientific research institutions, and enterprises. The maximum number of team members is 3.
- To ensure compliance with local regulations during the competition, all participants should comply with the export control laws of their country. In case of any negative impact on the competition due to violation of any export control laws, the team members reserve the right to disqualify the relevant contestants and take legal action.
- Participants shall express their interest to participate in this Grand Challenge by sending an email to the organizer Yongfu Li (yongfu.li@sjtu.edu.cn) and are invited to submit their work as BioCAS papers selecting the special session "Lung Sound Design Contest". The papers will be regularly reviewed and, if accepted, must be presented at BioCAS 2023.
Awards
- Total Prize of USD10,000.
- IEEE CASS Travel Student Grant (Up to USD2,500/Team).
- Top-4 team will be invited to present their work in IEEE BioCAS 2023.
- Top-2 team will get invited to submit their work to IEEE TBioCAS Special Issue.
Challenge and Dataset
Datasets
Our database is the first open access respiratory sound database in pediatric population, aging from 1 month to 18 years old. The respiratory sounds contained in the dataset were recorded at the pediatric respiratory department in Shanghai Children’s Medical Center (SCMC) using Yunting model II Stethoscope.
The recordings are saved in .wav format with naming rules as follows: Each name is compromised with 5 elements separated with underscores, including the patient number, age, gender, the recording location, and the recording number of the participants.
Patient number (e.g., 65101170)
Age (e.g., 0.4)
Gender
a. Male (0)
b. Female (1)
Recording location
a. left posterior (p1)
b. left lateral (p2)
c. right posterior (p3)
d. right lateral (p4)
Recording number (e.g., 3246)
The annotations at the record and event level are provided in this database. At the record level, each recording with poor signal quality was annotated as Poor Quality, while the recordings with high signal quality was annotated as Normal, CAS, DAS, or CAS & DAS according to the presence/absence of continuous/discontinuous adventitious respiratory sounds. At the event level, each recording was segmented into multiple respiratory events and annotated as Normal, Rhonchi, Wheeze, Stridor, Coarse Crackle, Fine Crackle, or Wheeze+Crackle.
The annotation information of each recording is saved in .json format with the same filename, which contains the annotation at record level and event level. The annotation at record level is Normal, CAS, DAS, CAS & DAS or Poor Quality. The annotation at event level consists of the start (ms) and the end (ms) of respiratory events, and the corresponding type of the respiratory events (Normal, Rhonchi, Wheeze, Stridor, Coarse Crackle, Fine Crackle, Wheeze+Crackle).
An example of the annotation file is as follow:
{
"recording_annotation": "Normal",
"event_annotation": [
{
"start": 342,
"end": 2515,
"type": "Normal"
}, {
"start": 2557,
"end": 3776,
"type": "Normal"
}, {
"start": 4547,
"end": 5651,
"type": "Normal"
}, {
"start": 6439,
"end": 8065,
"type": "Normal"
}, {
"start": 8363,
"end": 9201,
"type": "Normal"
}
]
}
This database is freely available for research and can be downloaded from https://github.com/SJTU-YONGFU-RESEARCH-GRP/SPRSound. Publications using this database should cite the data collection in order to identify the database:
Qing Zhang, "SPRSound: Open-Source SJTU Paediatric Respiratory Sound Database," [Online]. Available: https://github.com/SJTU-YONGFU-RESEARCH-GRP/Lung-Sound-Database/
Main Tasks
Task 1 (Respiratory Sound Classification at Event Level):
Task 1-1 is a binary class classification challenge (Normal and Adventitious).
Task 1-2 is a multiclass classification challenge (Normal (N), Rhonchi (R), Wheeze (W), Stridor (S), Coarse Crackle (CC), Fine Crackle (FC), Wheeze & Crackle (WC)).
Task 2 (Respiratory Sound Classification at Record Level):
Task 2-1 is a ternary class classification challenge (Normal, Adventitious, and Poor Quality records).
Task 2-2 is a multiclass clasification challenge (Normal (N), CAS (C), DAS (D), CAS & DAS (CD), or Poor Quality (PQ) records).
Evaluation Metrics
Submissions are evaluated based on the following metrics including sensitivity (SE), specificity (SP), average score (AS), and harmonic score (HS) are as follows:
Task 1:
(1)
(2)
(3)
(4)
Task 2:
(1)
(2)
(3)
(4)
Rules
Every challenge participant agrees to use the provided data only in the scope of the DATA USE AND CONFIDENTIALITY AGREEMENT for access to data.
For training the model, no external data is allowed except the official dataset provided in this challenge and pre-trained models using the ImageNet database such as VGG16, InceptionV3, ResNet, etc.
The challenge does not encourage excessive stacking of models and hardware to brush up the score of the challenge.
Every challenge member agrees that the decisions of the challenge committee will be final and binding on all matters related to this challenge. If there is any change to data, schedule, instructions of participation, or these rules, the registered participants will be notified of the email addresses they provided when they are registered.
If an unforeseen or unexpected event (including, but not limited to: someone cheating; a virus, bug, or catastrophic event corrupting data or the submission platform; someone discovering a flaw in the data or modalities of the challenge) that cannot be reasonably anticipated or controlled, (also referred to as force majeure) affects the fairness and/or integrity of this challenge, the committee reserve the right to cancel, change or suspend this challenge. This right is reserved whether the event is due to human or technical error.
Submission
All individuals, teams and each member should submit the data use agreement in advance. (If not, your submissions will not be accepted.)
Multiple submissions are allowed. Please limit your submission to 1 per day.
The submitted files must be compressed in zip format. The main script (main.py) should be provided in the submitted files, which is the executable file for model evaluation. The readme.md and requirements.txt should be provided to describe the model architecture and list all the dependencies of your python project, respectively.
The command line and parameter requirements of the execution code are as follows:
python3 main.py --task task_level --wav /path/to/wav_path/ --out /path/to/output.jsonThe task_level is 11, 12, 21, or 22 (representing task 1-1, task 1-2, task 2-1, and task 2-2, respectively).
Notice that the wav files for respiratory sound classification at event level (task 1-1 & task 1-2) are segmented wav files.
The format of output json (UTF-8) is as follows:
{
wav_file_name1: predicted_type1,
wav_file_name2: predicted_type2,
wav_file_name3: predicted_type3,
…
}Live Demo Submission
As part of this grand challenge, we encourage our participants to submit a live demo in which you deploy your model on a cell phone for real-time applications. We hope you can open source your project to help accelerate this research direction.
You can refer to our past demo (BioCAS 2019) in the following github link:
https://github.com/SJTU-YONGFU-RESEARCH-GRP/Lung-Sound-Classification-System-LungSys-I