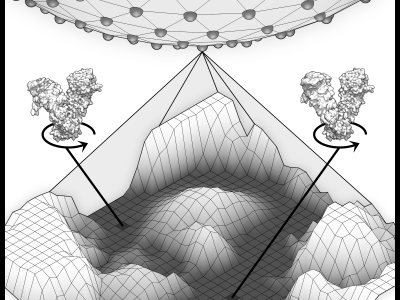
This repository contains code to apply the ESPER method to quasi-continuum models of biomolecules exhibiting multiple degrees of freedom, as described in Seitz et al. (2022, IEEE TCI). As inputs into ESPER, detailed instructions are also provided for generating custom synthetic datasets with increasing complexity to mirror known cryo-EM image attributes.
- Categories:

